Project: Automatic hyperparameter selection for t-SNE/UMAP
Description
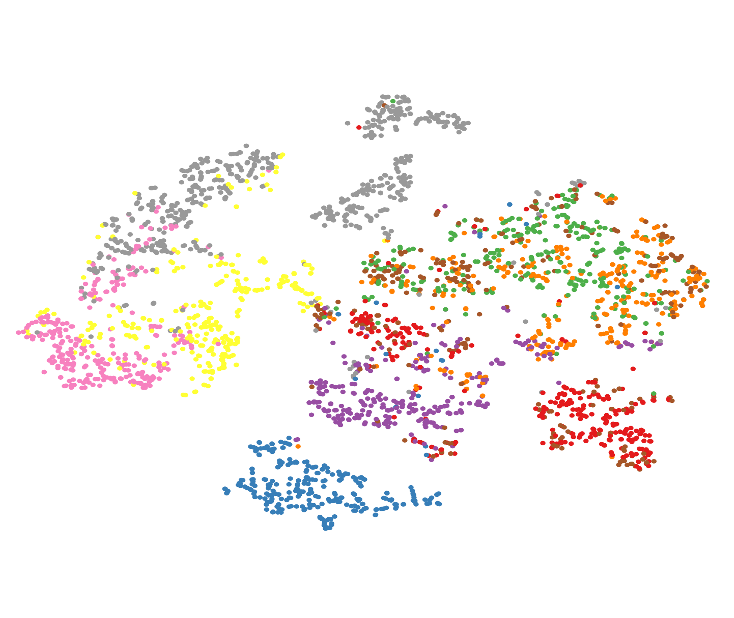
Local dimensionality reduction (DR) techniques such as t-SNE or UMAP are widely used in domains such as single-cell analysis, genome data, or protein folding to visualize and interpret complex, high-dimensional datasets. A key challenge for these techniques is setting the neighborhood parameters as they strongly influence the final projection created by the technique [1,2]. In t-SNE, this parameter is called perplexity, while UMAP uses n_neighbors. Choosing this value poorly can lead to misleading representations and incorrect interpretations of the data’s structure.
Although heuristics exist for setting these neighborhood parameters, they are often arbitrary and do not generalize well across different datasets. Existing approaches for automatic hyperparameter selection typically try to tune all hyperparameters of the technique simultaneously. However, the neighborhood size has a vastly different impact than the hyperparameters for optimization, making it difficult to define a suitable objective function.
In this project, we will create a novel approach to automatically set the neighborhood parameter of t-SNE and UMAP based on properties of the high-dimensional data.
[1] https://distill.pub/2016/misread-tsne/
[2] https://pair-code.github.io/understanding-umap/
Details
- Supervisor
-
 Diede van der Hoorn
Diede van der Hoorn
- Secondary supervisor
-
 Fernando Paulovich
Fernando Paulovich
- Interested?
- Get in contact