back to list
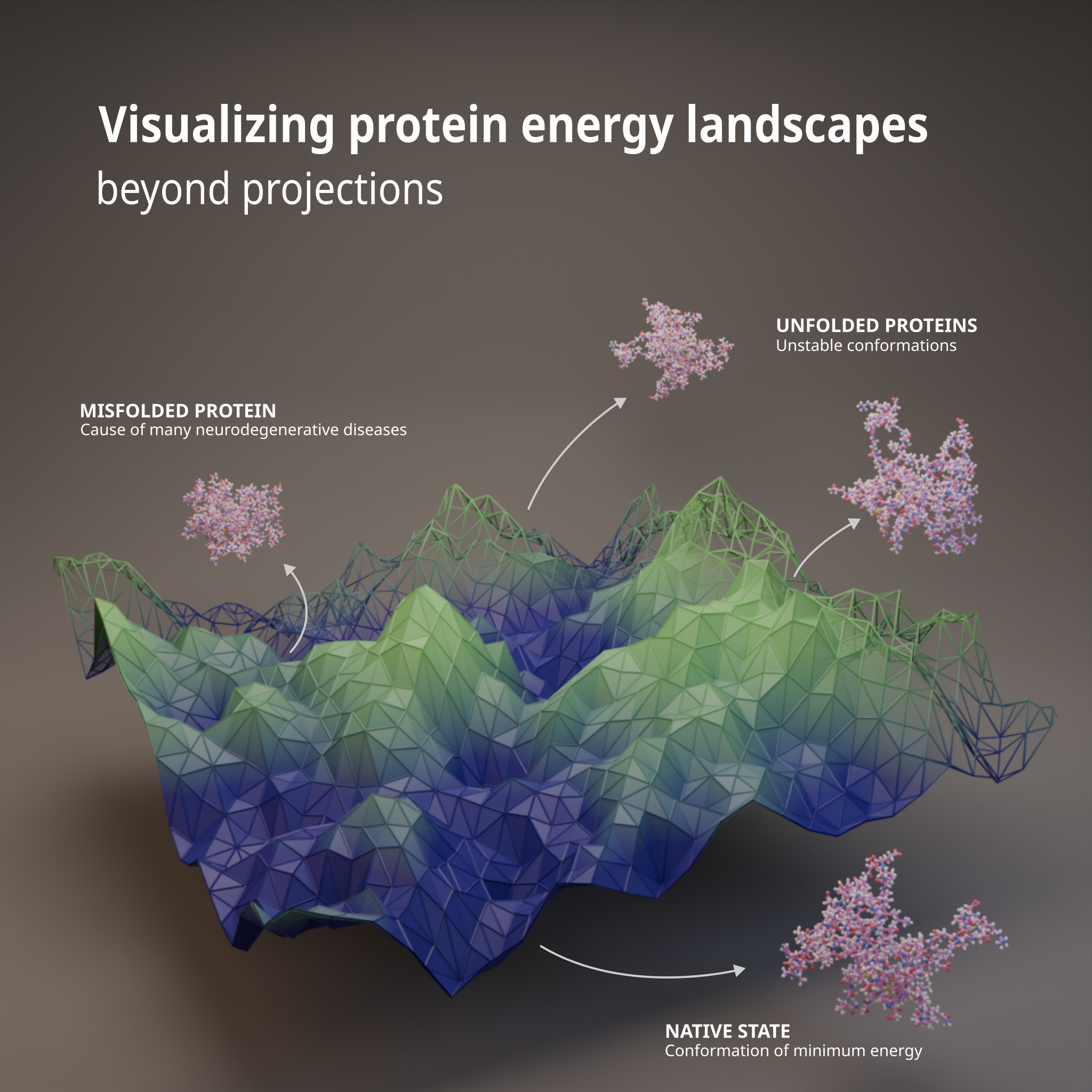
Project: Visualizing Protein Energy Landscapes Beyond Projections
Description

Protein folding is a complex, high‑dimensional dynamical process in which a protein transitions through a vast space of conformations toward energetically favorable folded states. This process is often conceptualized as an energy landscape, where stable structures correspond to minima, metastable states to local basins, and folding pathways to transitions between these regions.
To make this complexity interpretable, current approaches frequently rely on dimensionality‑reduction techniques—such as PCA, t‑SNE, UMAP, or diffusion maps—to project high‑dimensional simulation data into two or three dimensions. These low‑dimensional embeddings are then used to visualize folding pathways, clusters of conformations, and energy minima, as seen in recent work on energy‑landscape visualization for molecular dynamics simulations.
While dimensionality reduction has proven useful, it also introduces fundamental limitations. DR methods may distort distances, merge distinct folding pathways, obscure kinetic barriers, or exaggerate apparent clusters. As a result, visual interpretations of folding mechanisms, metastability, and transition states may depend more on the chosen projection than on the underlying physics of the folding process.
This master's project investigates how visual analytics techniques beyond standard dimensionality reduction can be developed to better represent and explore protein folding energy landscapes. The goal is not necessarily to discard DR entirely, but to replace, augment, or contextualize DR layouts with representations that preserve physically meaningful structures such as energy barriers, transition pathways, and kinetic relationships.
Research Questions
This project explores questions such as:
- What critical information about protein folding is lost or distorted by standard dimensionality‑reduction layouts?
- How can the energy landscape metaphor be visualized more faithfully without relying solely on low‑dimensional global embeddings?
- How can folding pathways, metastable basins, and transitions be represented in ways that remain interpretable to domain scientists?
- How can interactive visual analytics support hypothesis generation and exploration of folding mechanisms?
Approach
Data & Modeling
- Use molecular dynamics or protein folding simulation data containing:
- High‑dimensional conformational states,
- Associated energies or free‑energy estimates,
- Temporal or transition information.
- Use DR‑based embeddings as a baseline for comparison.
Visualization & Representation Design
The project may explore alternatives or complements to DR, such as:
- Graph‑based representations of conformational transitions and metastable states,
- Energy‑centric views that explicitly encode barriers and basins,
- Hybrid multi‑view systems combining local DR with global structural overviews,
- Topology‑aware or hierarchy‑based layouts to reveal folding funnels and branches,
- Interactive slicing or projection techniques guided by physically meaningful coordinates rather than abstract dimensions.
Expected Outcome
The expected outcome is a visual analytics prototype that demonstrates improved ways of exploring protein folding energy landscapes beyond traditional DR projections. The project will provide insights into how visualization choices influence scientific understanding and guidance for designing robust, interpretable visualizations of complex molecular processes.
Relevance
- J. Ros, A. Arleo, R. G. Viegas, V. B. P. Leite and F. V. Paulovich, Challenges and Opportunities for the Visualization of Protein Energy Landscapes, in IEEE Computer Graphics and Applications, vol. 45, no. 5, pp. 49-63, Sept.-Oct. 2025, doi: 10.1109/MCG.2025.3592983
- Rafael Giordano Viegas, Ingrid B. S. Martins, Murilo Nogueira Sanches, Antonio B. Oliveira Junior, Juliana B. de Camargo, Fernando V. Paulovich, and Vitor B. P. Leite. ELViM: Exploring Biomolecular Energy Landscapes through Multidimensional Visualization. Journal of Chemical Information and Modeling 2024 64 (8), 3443-3450. DOI: 10.1021/acs.jcim.4c00034
Details
- Supervisor
-
 Fernando Paulovich
Fernando Paulovich
- Secondary supervisor
-
 Jaume Ros
Jaume Ros
- Interested?
- Get in contact